PDF) Genetic diversity of pea (Pisum sativum L.) genotypes assessed by pedigree, morphological and molecular data | Snjažana Bolarić - Academia.edu
High-Throughput Development of SSR Markers from Pea (Pisum sativum L.) Based on Next Generation Sequencing of a Purified Chinese Commercial Variety | PLOS ONE
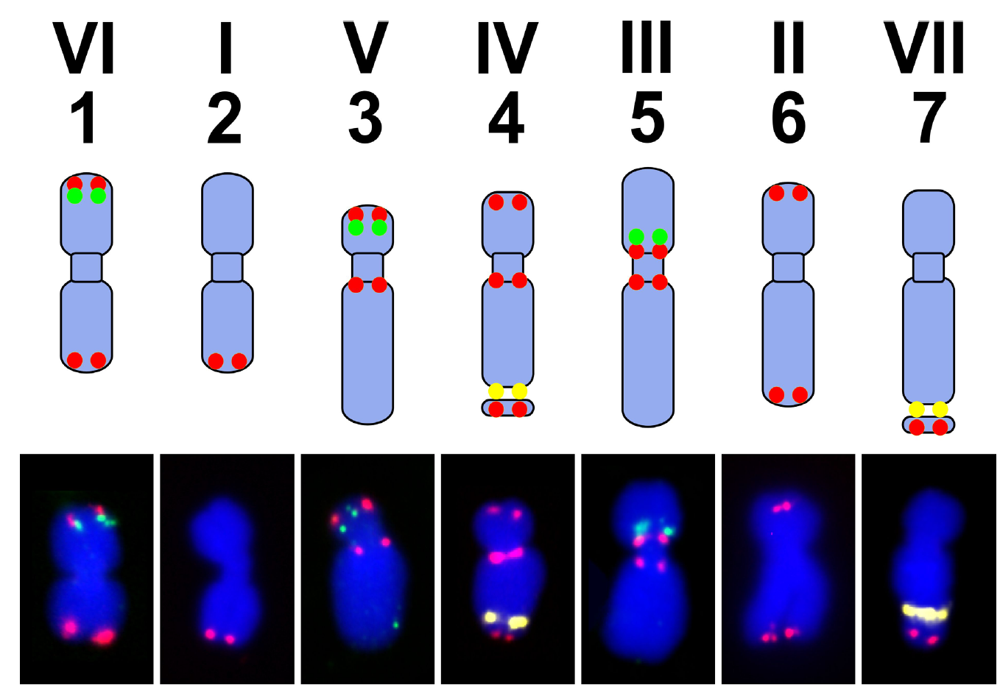
![PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6b50c024ad7293ae031ff7e23da4eb0630955b03/4-Figure2-1.png)
PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar
![PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6b50c024ad7293ae031ff7e23da4eb0630955b03/3-Table2-1.png)
PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar

PDF) Microsatellite markers for powdery mildew resistance in pea (Pisum sativum L.) | Stine Tuvesson - Academia.edu

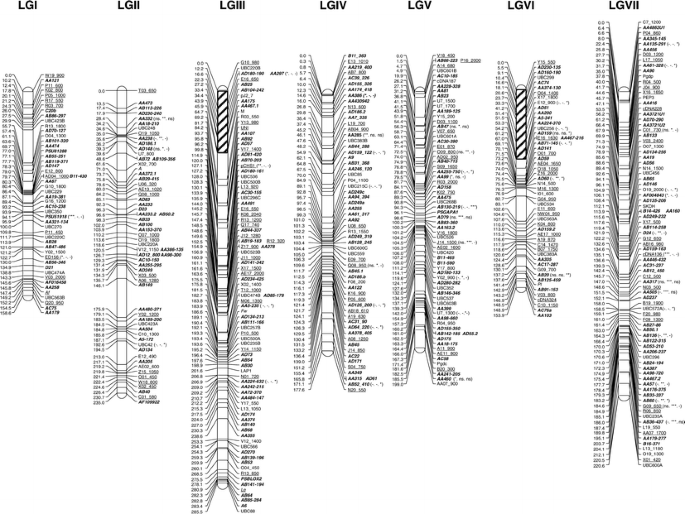
Genome-wide SNP identification, linkage map construction and QTL mapping for seed mineral concentrations and contents in pea (Pisum sativum L.) | BMC Plant Biology | Full Text
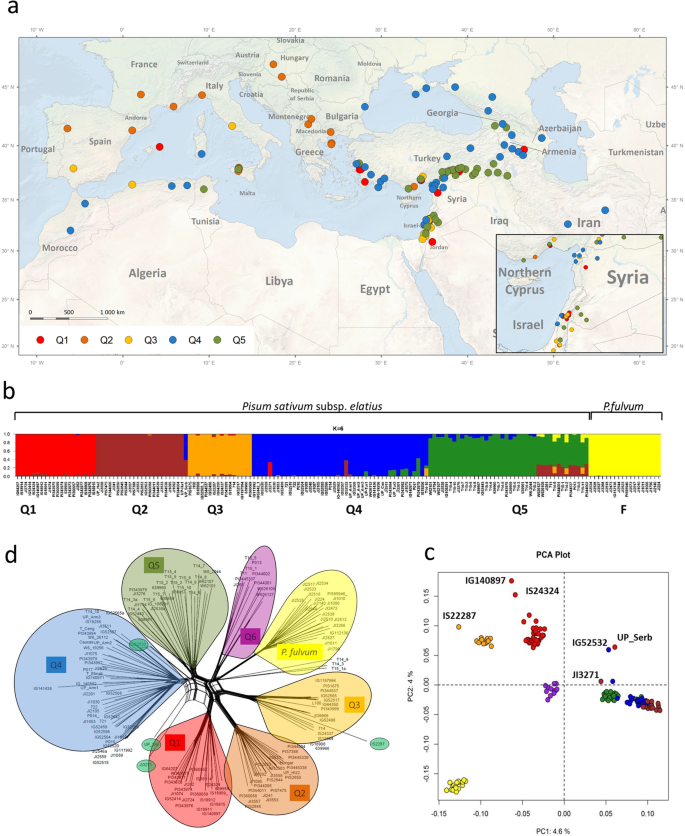
Genomic diversity and macroecology of the crop wild relatives of domesticated pea | Scientific Reports
Pea Marker Database (PMD) – A new online database combining known pea (Pisum sativum L.) gene-based markers | PLOS ONE

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect

A community resource for exploring and utilizing genetic diversity in the USDA pea single plant plus collection | Horticulture Research
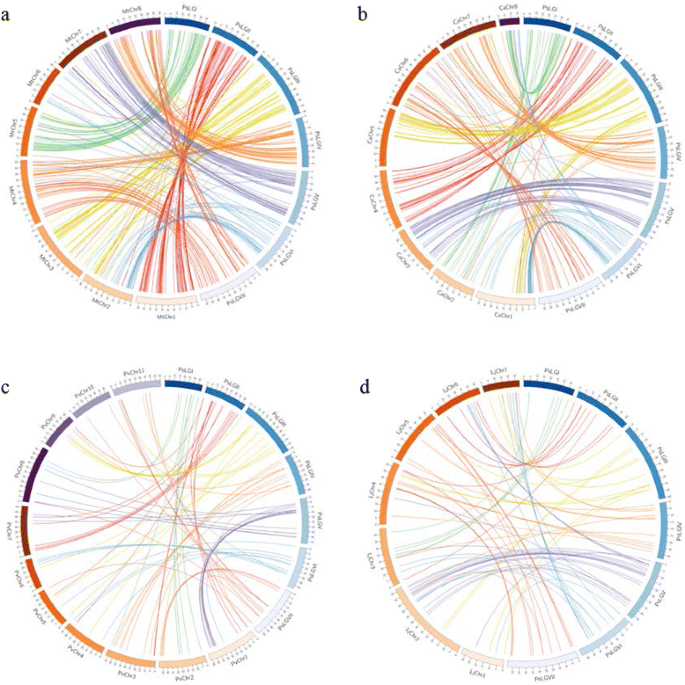
Genome-wide SNP identification, linkage map construction and QTL mapping for seed mineral concentrations and contents in pea (Pisum sativum L.) | BMC Plant Biology | Full Text
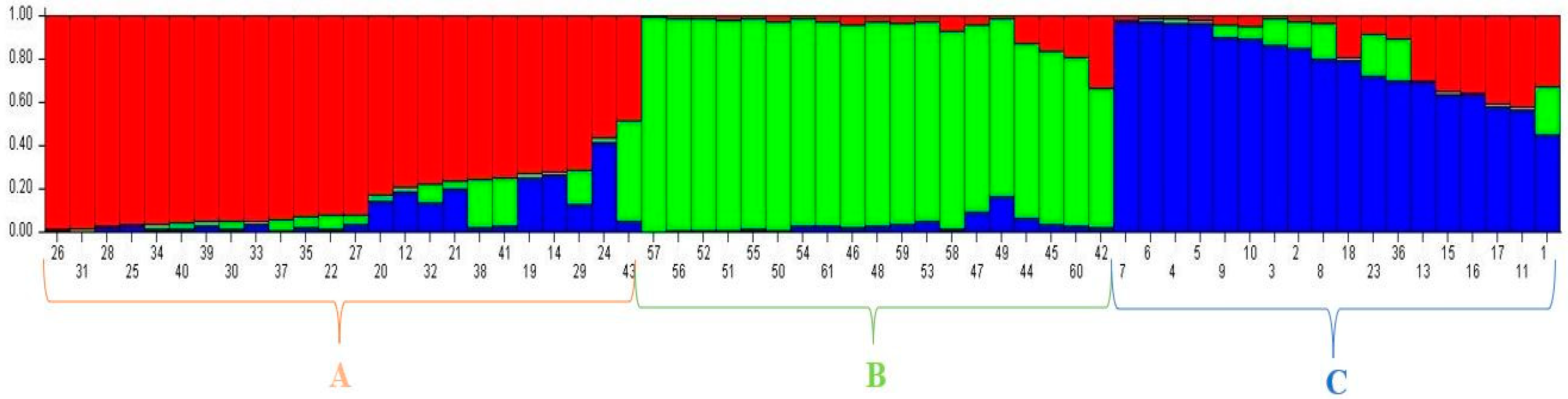
Genes | Free Full-Text | SSR-Based Molecular Identification and Population Structure Analysis for Forage Pea (Pisum sativum var. arvense L.) Landraces | HTML

PDF) Genetic Diversity and Population Structure Among Pea (Pisum sativum L.) Cultivars as Revealed by Simple Sequence Repeat and Novel Genic Markers

PDF) Estimation of pea ( Pisum sativum L.) microsatellite mutation rate based on pedigree and single-seed descent analyses | Petr Smykal - Academia.edu