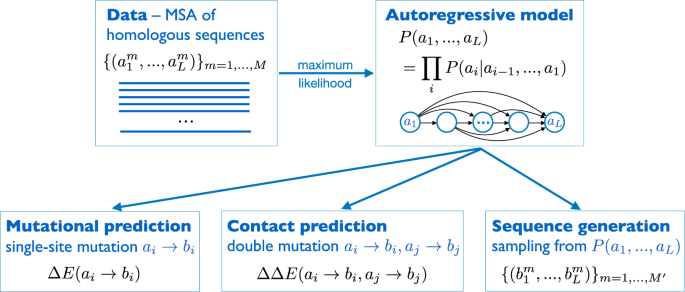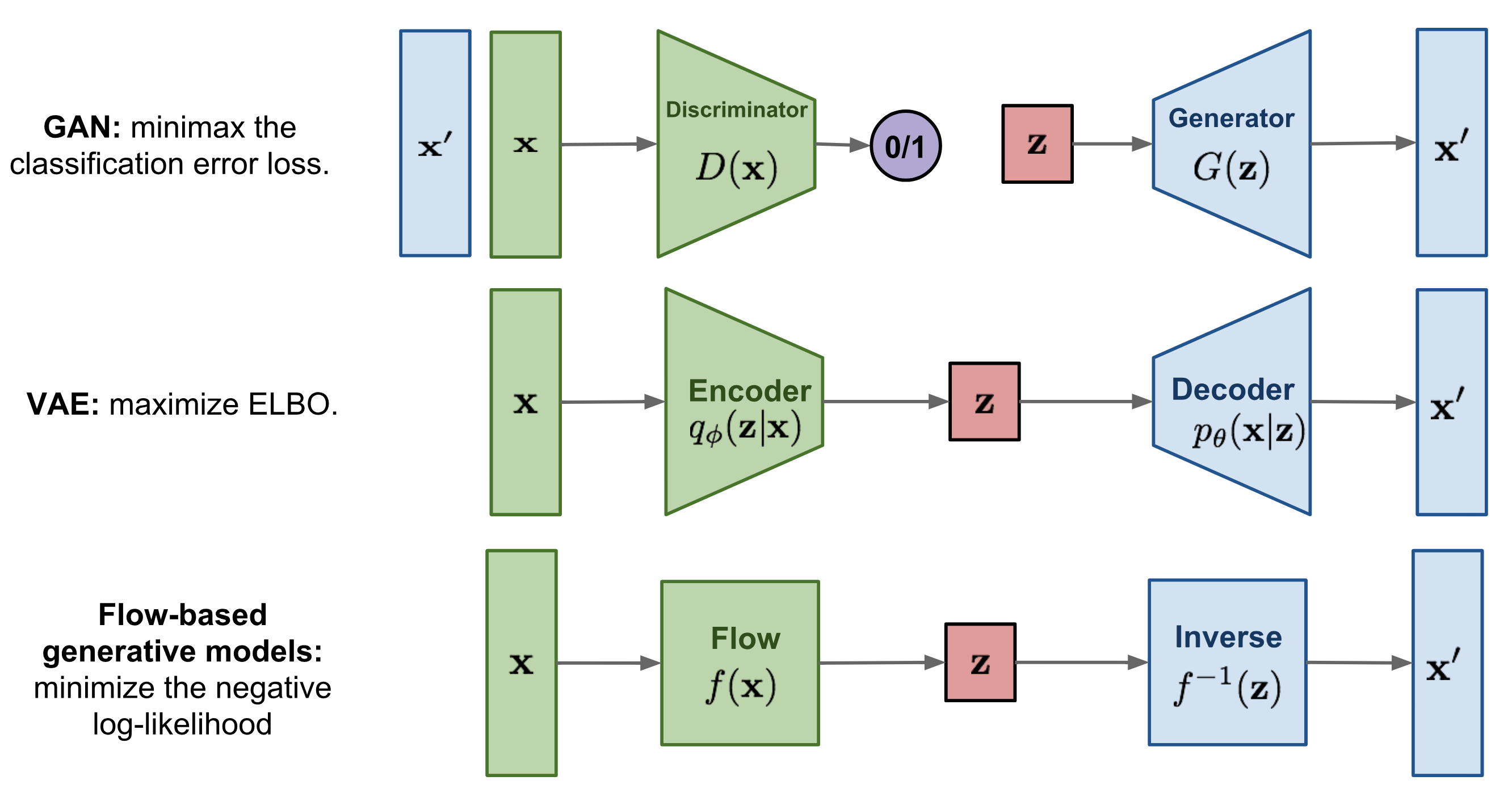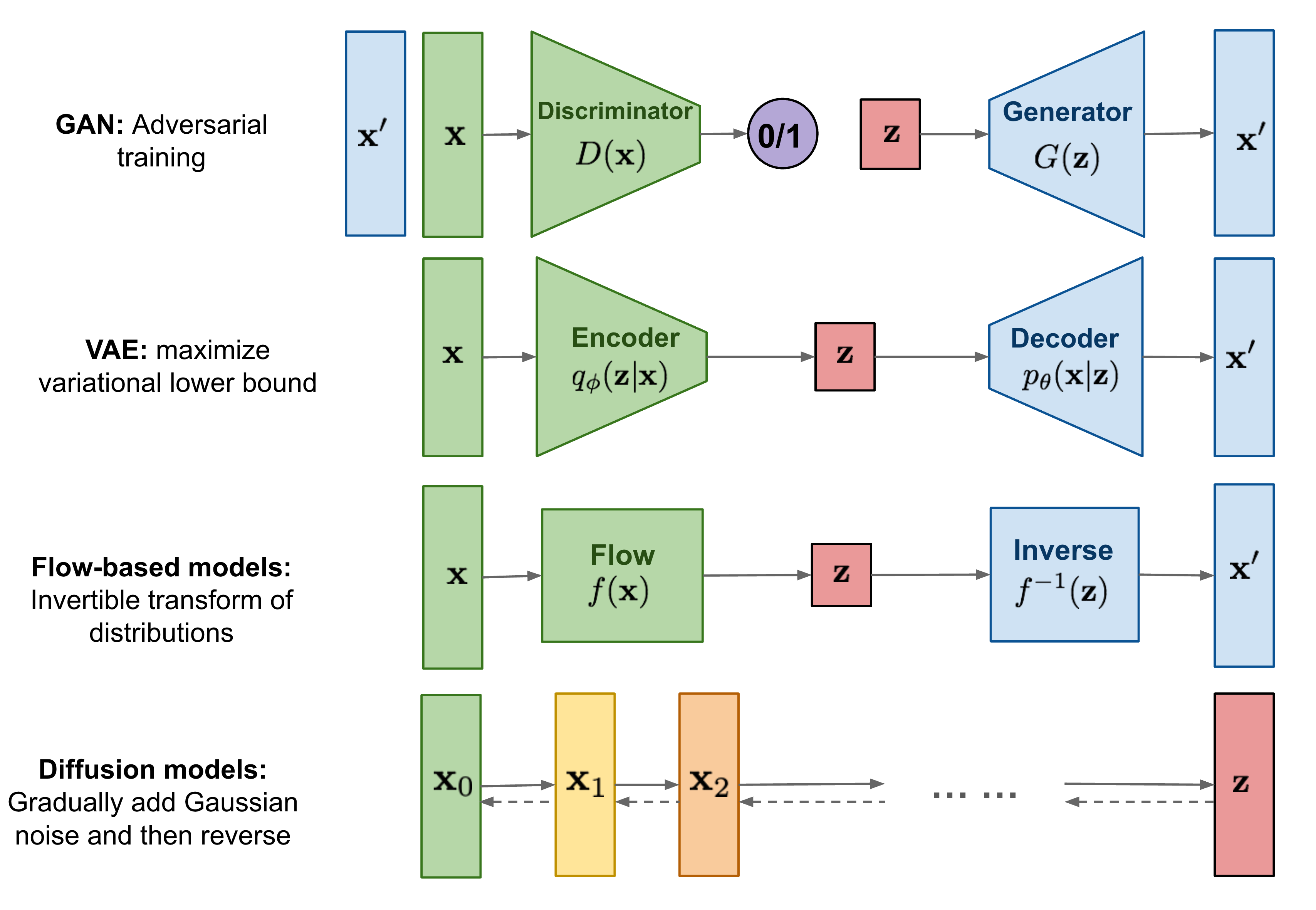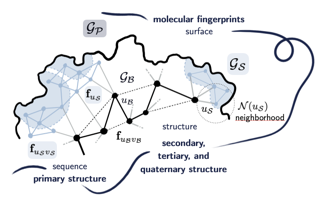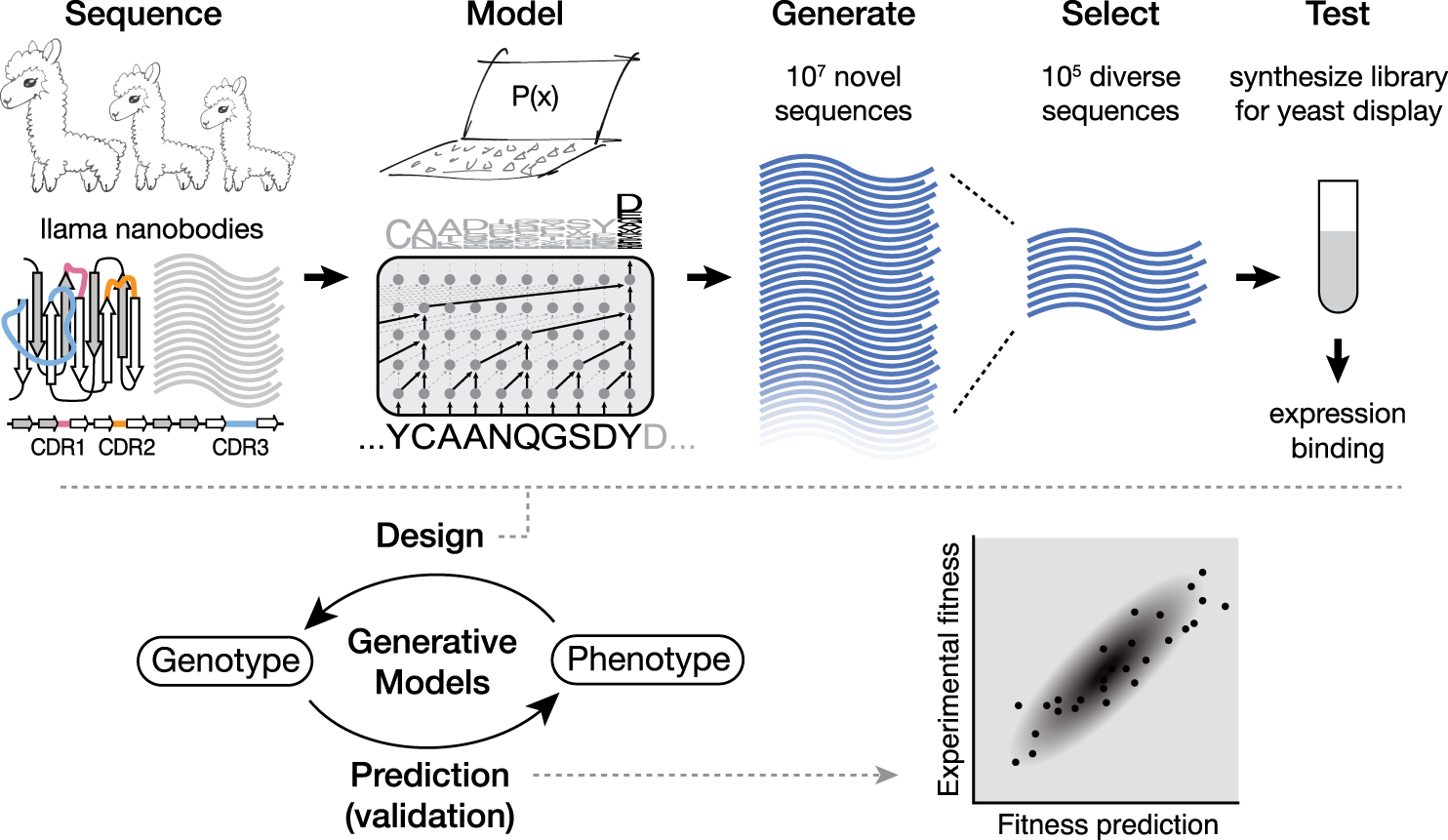
Protein design and variant prediction using autoregressive generative models | Nature Communications

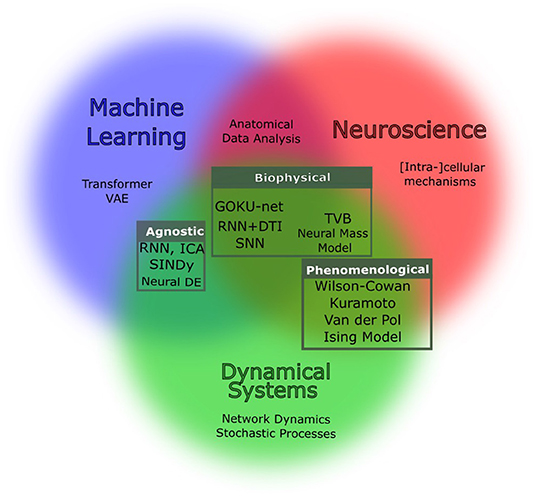
Enhancing scientific discoveries in molecular biology with deep generative models | Molecular Systems Biology
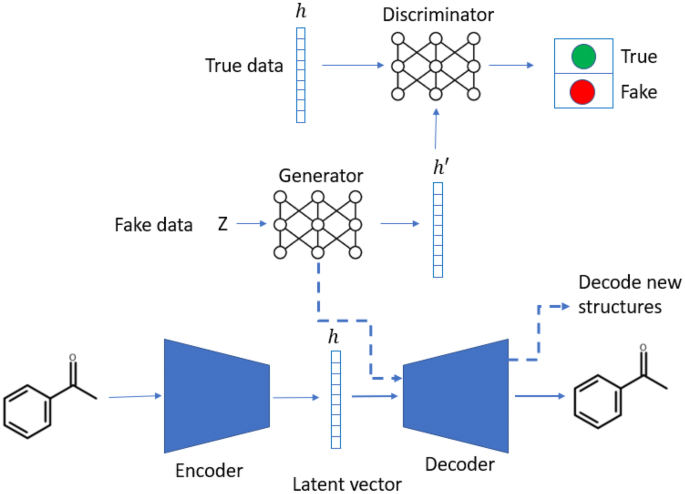
A de novo molecular generation method using latent vector based generative adversarial network | Journal of Cheminformatics | Full Text
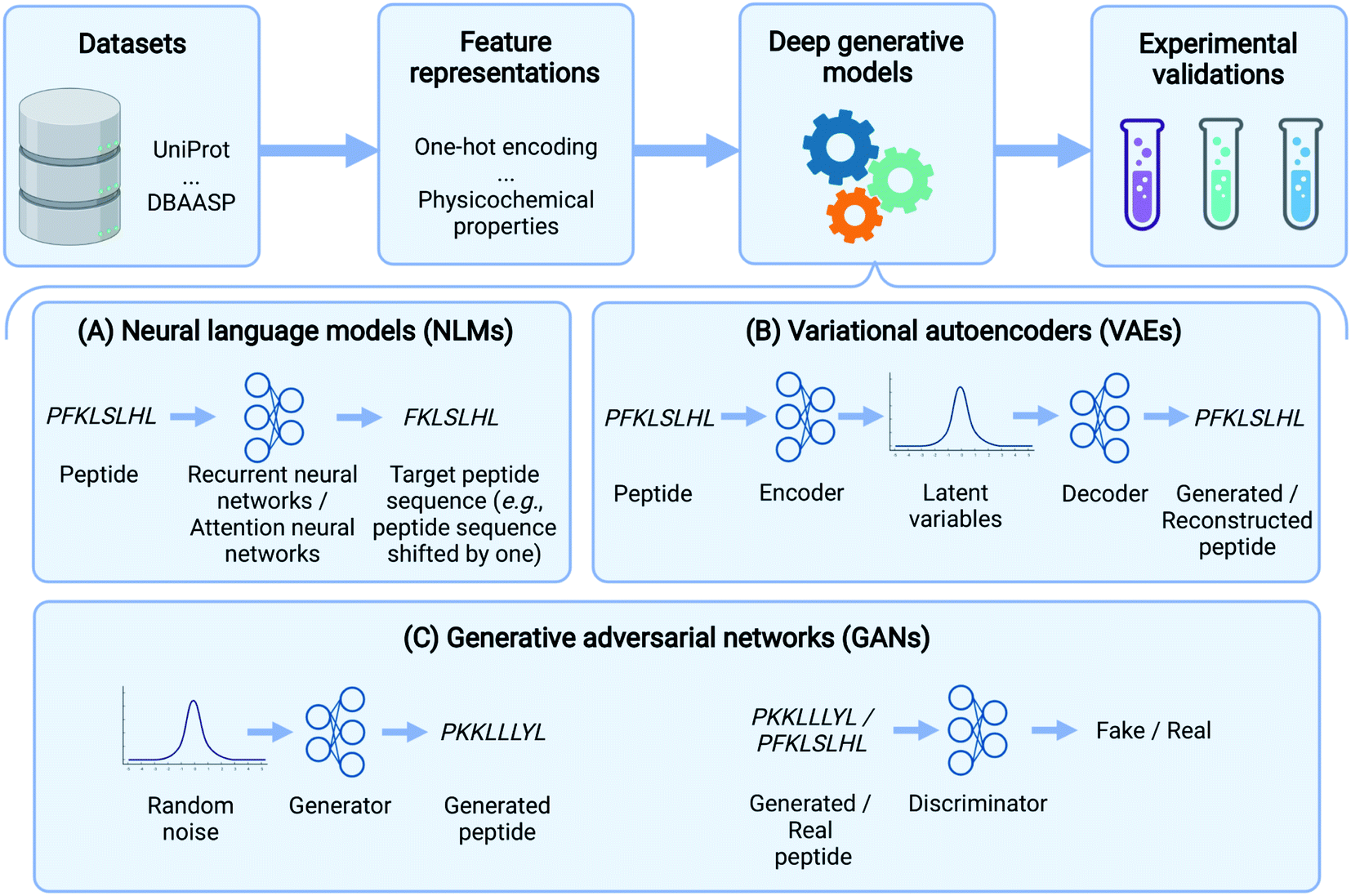
Deep generative models for peptide design - Digital Discovery (RSC Publishing) DOI:10.1039/D1DD00024A

Neural networks to learn protein sequence–function relationships from deep mutational scanning data | PNAS

A Generative Neural Network for Maximizing Fitness and Diversity of Synthetic DNA and Protein Sequences - ScienceDirect

Shape-Based Generative Modeling for de Novo Drug Design | Journal of Chemical Information and Modeling

Fold2Seq: A Joint Sequence(1D)-Fold(3D) Embedding-based Generative Model for Protein Design | DeepAI

SeqMed: Recommending Medication Combination with Sequence Generative Adversarial Nets | Semantic Scholar

From lattice-protein sequence space to inferred Potts model. Protein... | Download Scientific Diagram

Active Site Sequence Representations of Human Kinases Outperform Full Sequence Representations for Affinity Prediction and Inhibitor Generation: 3D Effects in a 1D Model | Journal of Chemical Information and Modeling

Generative probabilistic biological sequence models that account for mutational variability | bioRxiv